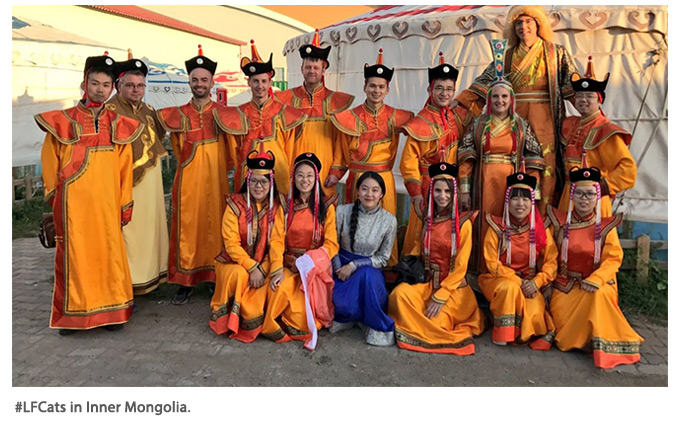
— By Sophien Kamoun with contributions from LFCats
On a cloudy Norwich day in 2011, post-docs Sebastian Schornack, Sylvain Raffaele, and Tolga Bozkurt were having a typical British lunch of fish and chips with mushy peas with their supervisor Sophien Kamoun. Somehow, the discussion turned to the importance of sustained productivity. Kamoun, in his usual hyperbolic style, pointed out that now that each one of them had just published notable papers (Schornack et al., 2010; Raffaele et al., 2010; Bozkurt et al., 2011), they should beware of not behaving like “lazy fat cats” and think hard about their next papers. Not everyone left the lunch in the happiest mood. One day later, after discussion with another post-doc, Mireille van Damme, Schornack and colleagues decided to found the Lazy Fat Cat Club (#LFCats). Schornack drafted a chart and was appointed as Chairman Féi māo (fat cat in Mandarin). The #LFCats ethos is that productive research requires a significant amount of communication and knowledge exchange, and informally discussing research is a perfect way of solving roadblocks and laying paths for the future. Casual meetings took place on a regular basis at The Sainsbury Laboratory, mainly on afternoon coffee breaks. The club continued to loosely grow and several other researchers joined the #LFCats. As the members moved on to start their own labs, the #LFCats “brand” helped nurture a lasting bond. Suomeng Dong, now a professor in the Department of Plant Pathology at Nanjing Agricultural University, coined the Chinese proverb “Fat cats cannot jump over the wall” to challenge the #LFCats to work collaboratively to solve problems and “jump over the wall.”

In this report, we summarize the key findings presented at the workshop.
The event kicked off with a keynote talk by
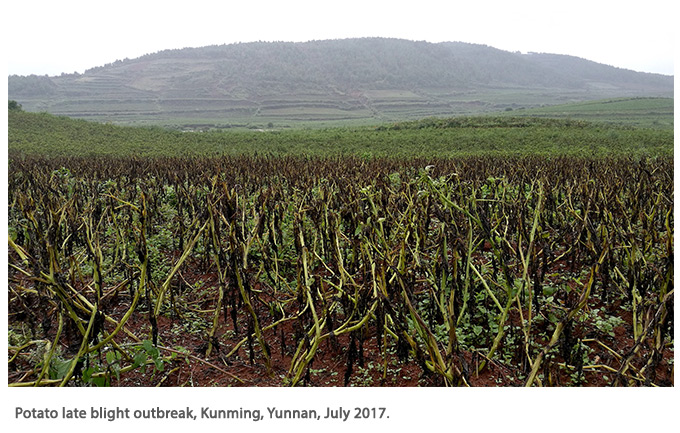
Yuanchao Wang, Nanjing Agricultural University. Wang reminded the audience of the importance of oomycete diseases in China by showing
dramatic photos of a potato late blight outbreak taken two weeks earlier in Kunming, Yunnan. Wang then described
elegant detective work to unravel one facet of the arms-race coevolution between Phytophthora and plants. The projects started with the identification of a secreted
Phytophthora sojae protein, named XEG1, that triggers hypersensitive cell death in
Nicotiana benthamiana. Using a gene-silencing loss-of-function screen, Wang and colleagues identified a receptor-like protein, RXEG1, that binds XEG1 and is required for XEG1-induced ROS burst and immunity. This screen also resulted in the identification of a receptor-like kinase protein, RFX1, that forms a complex with RXEG1 and is also required for XEG1-induced immunity. XEG1 encodes hydrolase activity, and loss-of-function mutants obtained by CRISPR/Cas9 knock-outs in
P. sojae revealed that XEG1 is required for full virulence on soybean via its catalytic activity. Interestingly, plants secrete a glucanase inhibitor, GIP1, that inhibits GIP1 and contributes to host resistance to
P. sojae. Remarkably, XEG1 has duplicated in
Phytophthora genomes, and a closely linked paralog, XLP1, present in head-to-head configuration with XEG1 in the
P. sojae genome, also binds soybean GIP1 but lacks catalytic activity. Binding of XLP1 to GIP1 is essential for virulence and competes with XEG1 for GIP1 binding, thus, counteracting GIP1 contribution to host defense. Thus,
P. sojae has evolved XLP1 as a decoy as a counter-defense strategy. Given that both XEG1 and XLP1 are widely distributed in
Phytophthora, the XLP1 decoy has evolved early in the genus, suggesting that GIP1-type defense response is a common anti-
Phytophthora defense response in plants. This works also highlights the importance of apoplastic interactions as a site of the antagonistic interactions between oomycetes and plants. Interestingly,
Phytophthora also delivers RXLR effectors into host cells that suppress RXEG1-mediated immunity, highlighting a recurrent theme throughout the workshop—the endless arms race between pathogens and their hosts.
Wenbo Ma, University of California-Riverside, was another distinguished speaker to join the workshop. She described her pioneering work on
Phytophthora effectors that suppress RNA silencing in plant hosts. These
Phytophthora suppressors of RNA silencing (PSRs) promote infection by targeting various host regulators of RNAi. One question is why the pathogen has evolved to suppress host RNAi. Ma reported exciting new results on how host-produced phased siRNAs (or phasiRNAs) target
Phytophthora genes, therefore, raising the hypothesis that host siRNAs traffic into the pathogen as a defense measure. Thus,
Phytophthora PSR effectors suppress the production of these siRNAs as a counter-defense strategy in another elegant example of the continuous arms race between
Phytophthora and host plants.
A second keynote speaker was
Jianmin Zhou, Chinese Academy of Sciences, who centered his presentation on
BIK1, a PBS1-like receptor like cytoplasmic kinase that negatively regulates immunity triggered by bacterial flagellin. Zhou reviewed the intricate dissection of the molecular mechanisms underpinning the activation of flagellin-triggered immunity in
Arabidopsis; most interestingly, his recent functional genomics analyses of
Arabidopsis RLCK family (46 genes) and the MAPKKK family (60 genes). This enabled him to discover that several RLCKs and MAPKKKs are involved in responses triggered by other PRRs, notably following elicitation by fungal chitin. Redundancy is the bane of genetic analysis. Zhou discussed how CRISPR/Cas9 can be used to generate multiple loss-of-function mutants to study how RLCKs cooperate to modulate plant immunity and enable a robust response of the plant to environmental perturbation. The topic of genetic redundancy, and its importance in regulating plant-pathogen interactions, was another common theme of the workshop. It is gratifying that genetic analyses have started to unravel complex network-type interactions in plant-pathogen systems.
Chih-Hang Wu, The Sainsbury Laboratory, described his work on
a helper-sensor NLR immune network that also illustrates the importance of genetic redundancy in plant immunity. This NLR network includes several well-known sensor NLRs, which confer disease resistance to diverse plant pathogens, and a few helper NLRs that display varying levels of functional redundancy and specificity to different sensor NLRs. Phylogenetic analyses of plant NLR proteins revealed that this NLR network originated from an ancestral NLR pair that emerged prior to the split between caryophyllale and asterid plants. Wu also discussed how this NLR network contributes to evolvability and robustness of plant NLR-triggered immunity.
Rosa Lozano-Duran, Shanghai Center for Plant Stress Biology, started
her talk by
reminding us how virus proteins can be multifunctional—a lesson worth remembering for those who work on bacteria and eukaryotic pathogens. One of the challenges that viruses face is to move from cell to cell, a feat they achieve through the plasmodesmata—membrane structures that serve as cytoplasmic bridges between cells. Remarkably, antiviral RNA interference (RNAi) can also move from cell to cell in plants, allowing for systemic spread of this immune response. Lozano-Duran reported that the
C4 protein of the geminivirus Tomato yellow leaf curl virus inhibits the intercellular spread of RNAi. C4 interacts with the plant protein receptor-like kinase (RLK) BARELY ANY MERISTEM 1 (BAM1) and colocalizes with this protein at plasmodesmata. BAM1 and its closest paralogue, BAM2, function as positive regulators of the cell-to-cell movement of RNAi. This work is another illustration of how pathogens have coopted host processes to favor colonization and infection.
The topic of genome-wide pathogen response to environmental cues was also covered by
Kentaro Yoshida, Kobe University. Yoshida set up an ambitious experiment to monitor pathogen gene expression dynamics in field over time, ranging from daily to seasonal scales. He showed intriguing seasonal dynamics in pathogen gene expression, e.g., reporting how a set of genes are up-regulated after increases in ambient temperature or after host flowering. Yoshida also reported periodical patterns of gene expression, particularly of effector genes.
Suomeng Dong, Nanjing Agricultural University, may have the key to global patterns of gene expression regulation. For a long time, the molecular basis of DNA methylation in
Phytophthora remained mysterious and the classic eukaryotic 5-methylcytosine (5mC) modification could not be reliably detected. Instead, Dong described how N6-methyldeoxyadenine (6mA) is widespread in
Phytophthora and appears to modulate gene expression levels. Interestingly, unlike most eukaryotes, 6m adenine methyltransferases have duplicated and functional diversified in
Phytophthora, possibly as an adaptation to the repeat-driven genome expansion.